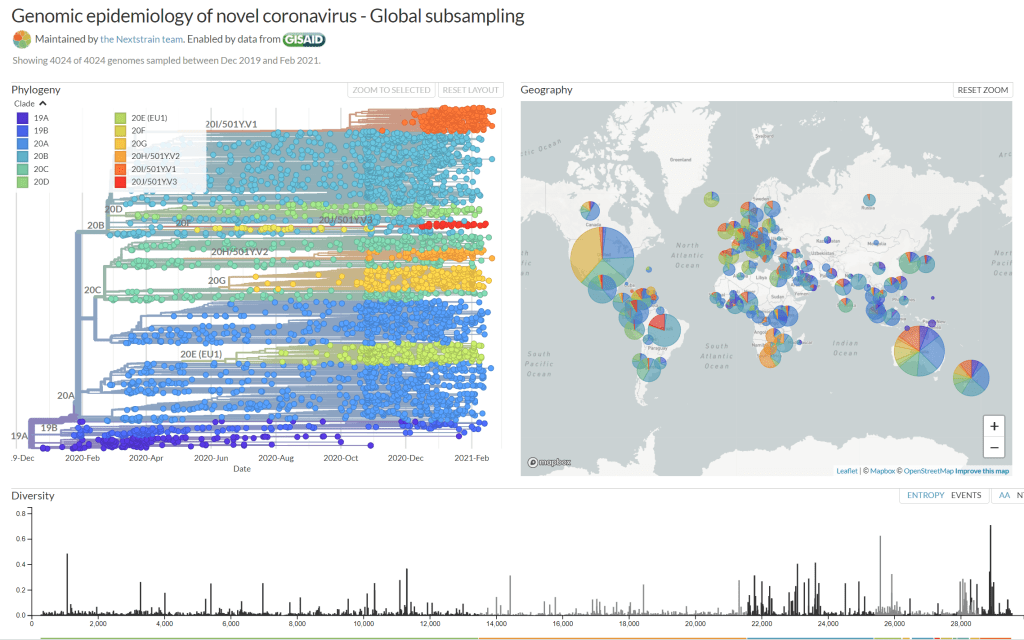
NextStrain update: https://nextstrain.org/ncov/global
For the longest time, CCP/CoVid-19 has shown remarkable viral genetic stability – remarkable for a zoonotic pathogen that only “recently jumped” species.
Even more remarkable in that the “jump” occurred without any known species as the vector.
And further remarkable that CCP/CoVid-19 is ~96% similar to a virus “discovered” in an abandoned mine in 2013, and only mentioned in 2020 but which no known samples are available (Zhou et al. (2020)).
This stability is particularly remarkable since CCP/CoVid-19 is a single-strand RNA virus – a class of viruses that normally evolve rapidly.
Only after widespread disperson did this stability finally begin to give way to evolution. Note the emergence of the new variants in late Q4, 2020 (upper right of the phylogeny diagram). The sudden ruse in reported COVID-19 cases are attributed to new SARS-CoV-2 variants 501Y.V1 (B.1.1.7) in the UK and 501Y.V2 (B.1.351) in South Africa.
These variants share a key mutation N501Y on the receptor binding domain (RBD) that make them more infectious to humans.
As reported by Tian et al. (2021), RBD N501Y mutations are of higher binding affinity to ACE2 than the wild type, a faster association rate, and slower dissociation rate.
Steered Molecular Dynamics (SMD) simulations on the dissociation of RBD-ACE2 complexes revealed that the N501Y introduced additional π-π and π-cation interaction for the higher force/interaction.
The reinforced interaction from N501Y mutation in RBD should play an essential role in the higher transmission of COVID-19 variants observed to date and likely to continue.
All in, quite “remarkable.”
Reference:
Tian, F., Tong, B., Sun, L., Shi, S., Zheng, B., Wang, Z., … Zheng, P. (2021). Mutation N501Y in RBD of spike protein strengthens the interaction between COVID-19 and its receptor ACE2 (p. 2021.02.14.431117). doi:10.1101/2021.02.14.431117
Zhou, P., Yang, X.-L., Wang, X.-G., Hu, B., Zhang, L., Zhang, W., … Shi, Z.-L. (2020). A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature, 579(7798), 270–273.