This week’s update.
Let’s also note the preprint released by Wells et al. regarding the evolutionary history of ACE2 usage.
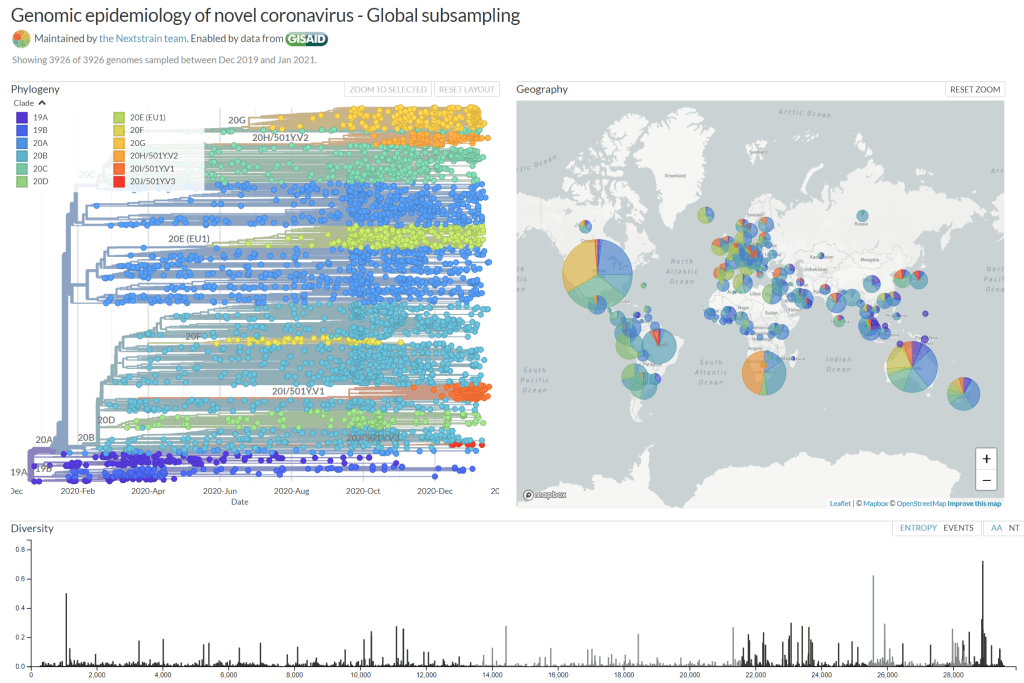
As evident in the Nexstrain phyologeny, CoVid-2/CCP (CoV-2) is a rather stable virus – unlike other zoonotics that have jumped species.
Wells examines possible genealogical histories for CoV-2 sequences. They demonstrate the lineage containing CoV-2 is most likely the ancestral ACE2-using lineage, and that recombination with at least one virus from this group conferred ACE2 usage to the lineage including SARS-CoV-1 at some time in the past. They propose a competitive release hypothesis to explain how this recombination event could have occurred and why it is evolutionarily advantageous.
This study also underscores the need for increased surveillance for sarbecoviruses in southwestern China, where most ACE2-using viruses have been found to date, as well as other regions such as Africa, where these viruses have only recently been discovered.
Importantly, genetic recombination has the potential to rapidly change the properties of a viral pathogen, and its presence is a crucial factor to consider in the development of treatments and vaccines.
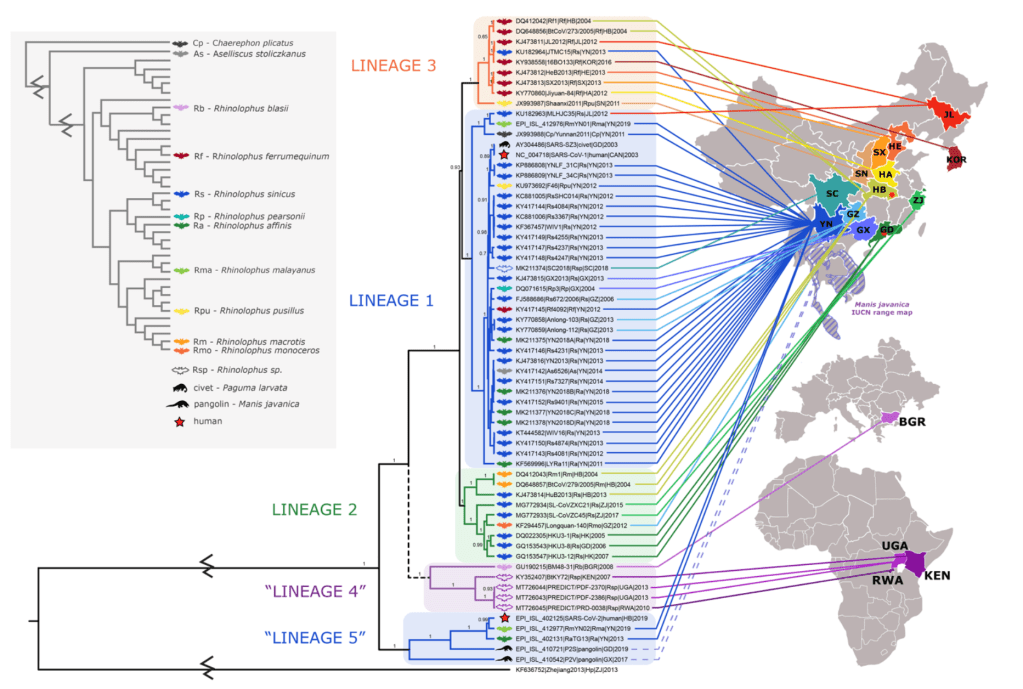
This finding of overlapping Lineage 1 and Lineage 5 viruses in geographic space is inconsistent with the previously observed pattern of biogeography for sarbecoviruses. SARS-CoV-2 was isolated first frompeople in Hubei Province and one of the pangolin viruses was isolated from an animal sampled in Guangdong, neither of which are Lineage 1 provinces. However, the true geographic origins of these viruses are unknown as it is possible they were anthropogenically transported to the regions in which they were detected. For example, the Malayan pangolin (Manis javanica) has a natural range that reaches southwestern China (Yunnan Province) at its northernmost edge and extends further south into Myanmar, Lao PDR, Thailand, and Vietnam. So, if they were naturally infected (as opposed to infection via wildlife trade), the infection was potentially not acquired from Guangdong Province. Similarly, SARS117 CoV-2 cannot be guaranteed to have emerged from bats in Hubei Province, as humans are highly mobile and the exact spillover event was not observed. If the clade containing SARS-CoV-2 and its close relatives is indeed endemic in animals in Yunnan and the nearby Southeast Asian regions as suggested by the presence of RaTG13, RmYN02, and the natural range of the Malayan pangolin, whatever mechanism is facilitating the biogeographical concordance of Lineages 1, 2 and 3 within China appears to no longer apply for the biogeography of Lineage 5, since they all appear to overlap in and around Yunnan Province.
Further, investigations into determinants of pathogenicity and transmission for CoVs and the genomic signatures of such features will be an important step towards the prediction of viruses with spillover potential, and distinguishing those with pandemic potential.
Finally, these findings reiterate the importance of recombination as a driver of spillover and emergence, particularly in the spike gene. If SARS-CoV-1 gained the ability to use hACE2 through recombination, other non-ACE2-using viruses could become human health threats through recombination as well. We know that recombination occurs much more frequently than just this single event with SARS-CoV-1, as the RdRp phylogeny does not mirror host phylogeny and the RBD tree has significantly different topology across all geographic lineages. In addition the bat virus RmYN02 appears to be recombinant in the opposite direction (Lineage 5 backbone with Lineage 1 RBD) [36], again supporting the hypothesis that recombination occurs between these lineages. Our analyses support two hypotheses: first, that sarbecoviruses frequently undergo recombination in this region of the genome, resulting in this pattern, and second, that sarbecoviruses are commonly shared amongst multiple host species, resulting in a lack of concordance with host species phylogeny and a reasonable opportunity for coinfection and recombination. Bats within the family Rhinolophidae have also repeatedly shown evidence of introgression between species [67–72], supporting the hypothesis that many species in this family have close contact with one another which may facilitate viral host switching. Given that we have shown that ACE2-using viruses are co-occurring with a large diversity of non-ACE2 using viruses in Yunnan available under aCC-BY-NC-ND 4.0 International license. Province and in a similar host landscape, recombination poses a significant threat to the emergence of novel sarbecoviruses.
Reference
Wells, H. L., Letko, M., Lasso, G., Ssebide, B., Nziza, J., Byarugaba, D. K., … Anthony, S. J. (2020). The evolutionary history of ACE2 usage within the coronavirus subgenus Sarbecovirus. BioRxiv.Org: The Preprint Server for Biology. doi:10.1101/2020.07.07.190546