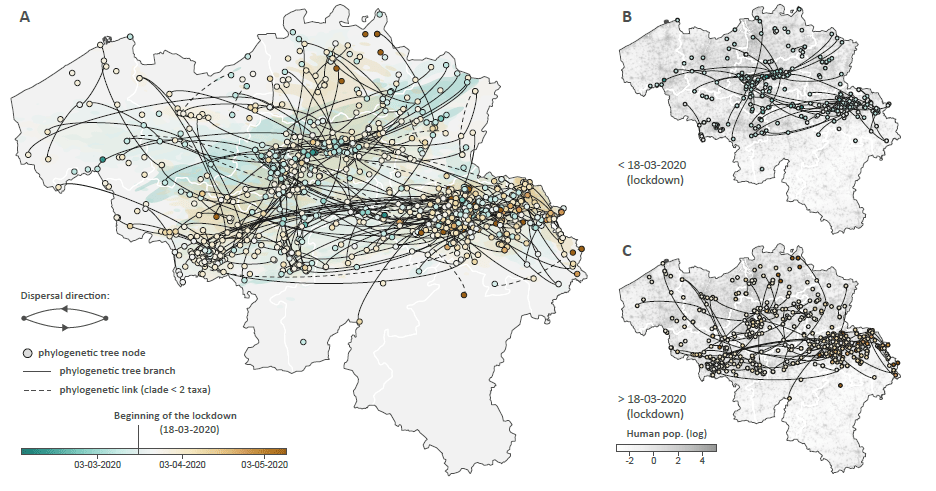
In a bioRxiv preprint first posted October 21, 2020 (and later published in Molecular Biology and Evolution), Dellacour et al. (2020) report their studies of the spatial density of available SARS-CoV-2 genomes mapped in the Belgium epidemic.
Their phylodynamic analysis demonstrates real-time dispersion dynamics of SARS-CoV-2 lineages.
Among other things, their spatially-explicit phylogeographic analyses highlight that the national lockdown had a relatively low impact on both the lineage dispersal velocity and long-distance dispersal events within Belgium.
Simply put, the Belgian lockdowns show no statistical evidence of materially impacting pandemic dispersion.

Reference
Simon Dellicour, Keith Durkin, Samuel L. Hong, Bert Vanmechelen, Joan Martí-Carreras, Mandev S. Gill, Cécile Meex, Sébastien Bontems, Emmanuel André, Marius Gilbert, Conor Walker, Nicola De Maio, Nuno R. Faria, James Hadfield, Marie-Pierre Hayette, Vincent Bours, Tony Wawina-Bokalanga, Maria Artesi, Guy Baele, Piet Maes (2020) A phylodynamic workflow to rapidly gain insights into the dispersal history and dynamics of SARS-CoV-2 lineages bioRxiv 2020.05.05.078758; doi: https://doi.org/10.1101/2020.05.05.078758
Now published in Molecular Biology and Evolution doi: 10.1093/molbev/msaa284